Simultaneous Inference of Past Demography and Selection from the Ancestral Recombination Graph under the Beta Coalescent
The reproductive mechanism of a species is a key driver of genome evolution. The standard Wright-Fisher model for the reproduction of individuals in a population assumes that each individual produces a number of offspring negligible compared to the total population size. Yet many species of plants, invertebrates, prokaryotes or fish exhibit neutrally skewed offspring distribution or strong selection events yielding few individuals to produce a number of offspring of up to the same magnitude as the population size. As a result, the genealogy of a sample is characterized by multiple individuals coalescing simultaneously to the same common ancestor. The current methods developed to detect such multiple merger events do not account for complex demographic scenarios or recombination, and require large sample sizes.
We tackle these limitations by developing two novel and different approaches to infer multiple merger events from sequence data or the ancestral recombination graph (ARG): a sequentially Markovian coalescent and a graph neural network (GNNcoal).
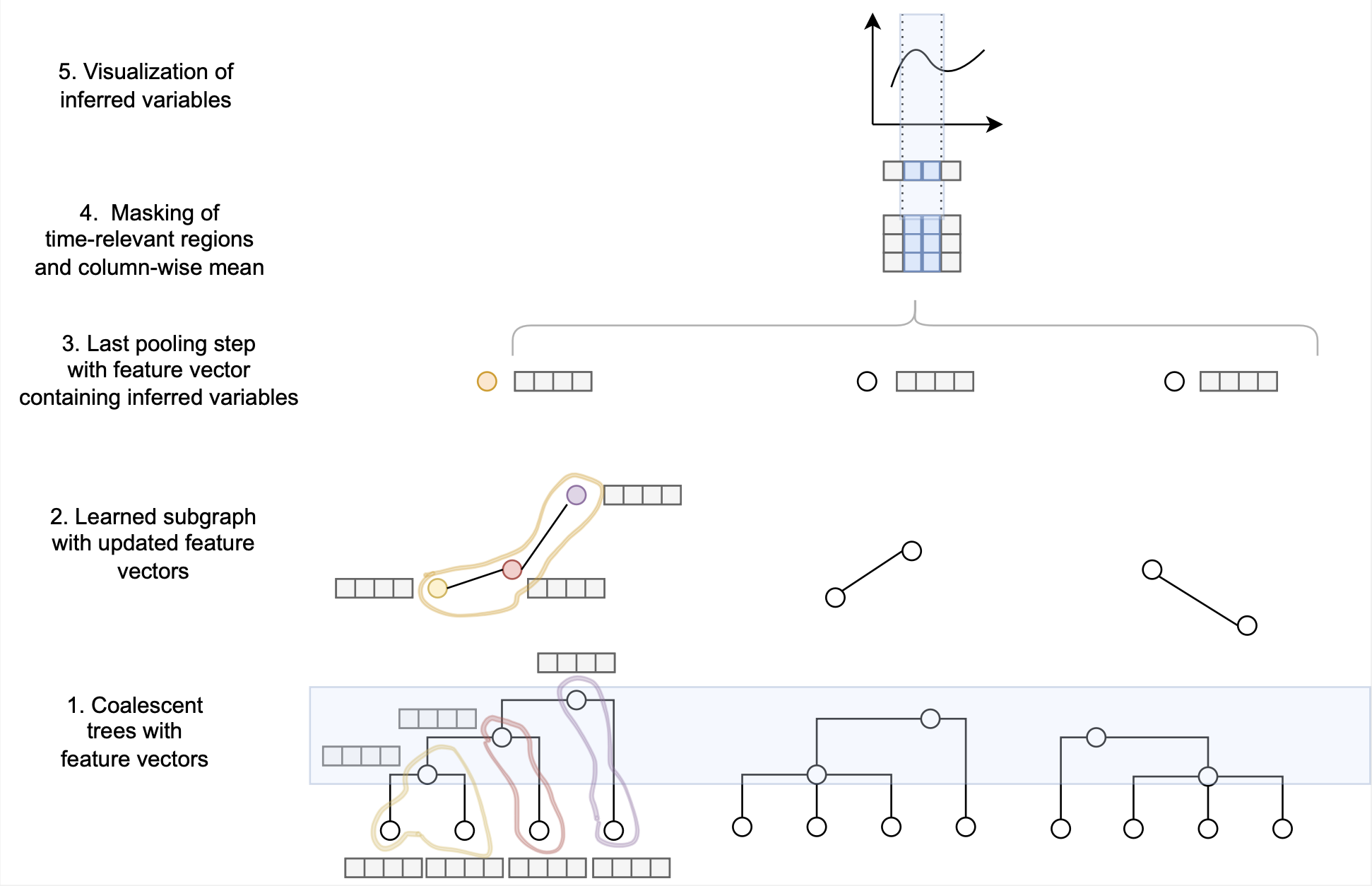
We first give proof of the accuracy of our methods to estimate the multiple merger parameter and past demographic history using simulated data under the beta-coalescent model. Secondly, we show that our approaches can also recover the effect of positive selective sweeps along the genome. Finally, we are able to distinguish skewed offspring distribution from selection while simultaneously inferring the past variation of population size.
Our findings stress the aptitude of neural networks to leverage information from the ARG for inference but also the urgent need for more accurate ARG inference approaches.
The paper is available here.